In 2019, Dr. Read and Dr. Hristrova went to the Anza Borrego Desert in California and discovered a desiccated stream in the San Felipe Creek of the Anza Borrego Desert. There they collected samples of desiccated cyanobacteria. Clarivel (my lab partner) and I were given the project to identify what species of cyanobacteria it is.
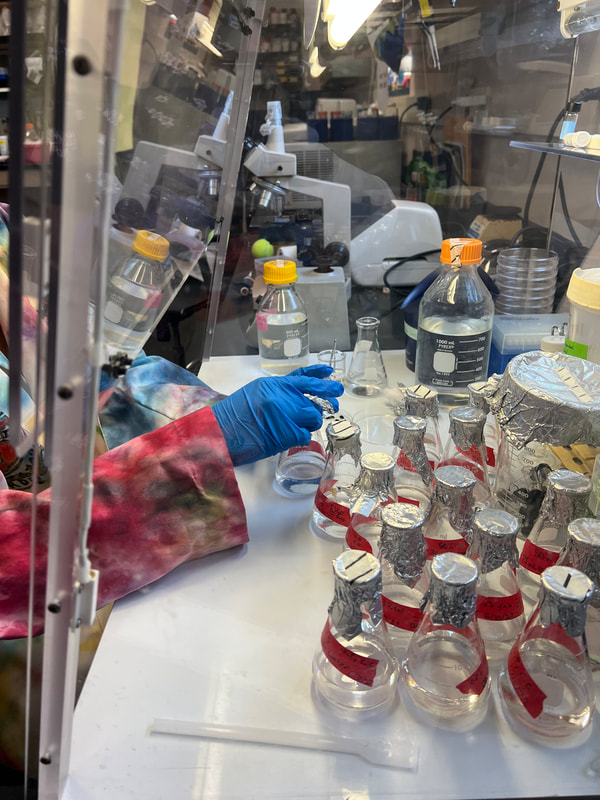
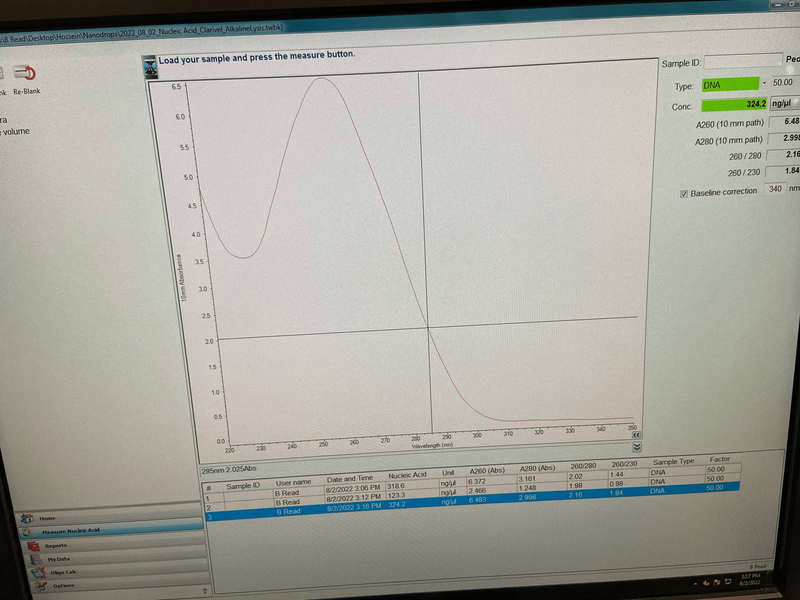
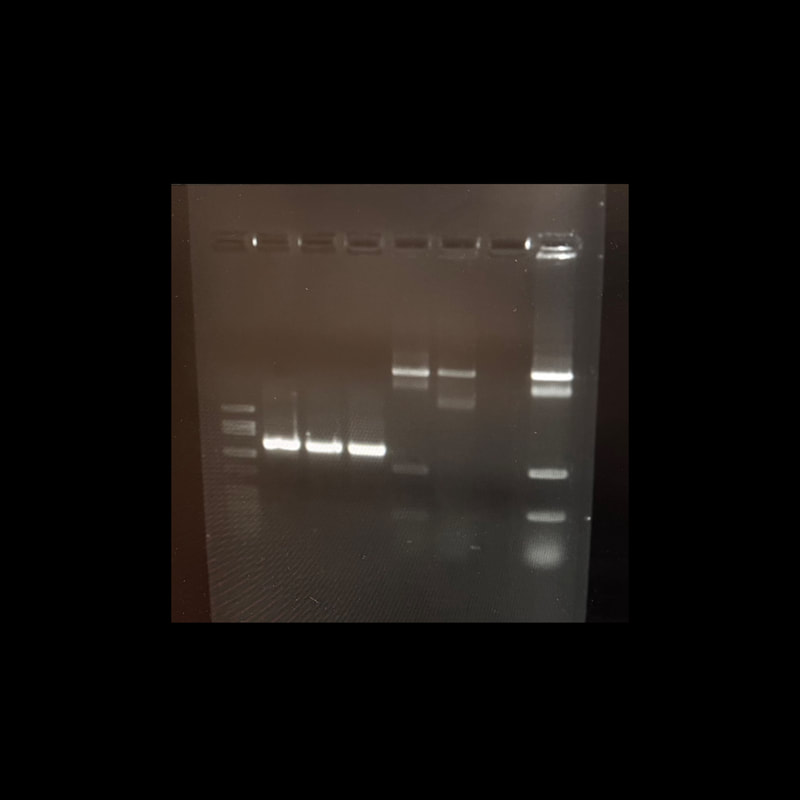
For the past 8 weeks, our initial goal for our project is to identify the unknown cyanobacterium from the Anza Borrego San Felipe Creek with whole genome sequencing and then to interrogate its phylogenetic history. During this time I have learned a lot of bioinformatic tools and techniques. I’ve also heard the horrors and tragedies of bioinformatics, and how important it is to maintain detailed notes of the purpose and methods used in computational analysis. With the help of Dr. Read, I was able to quickly get the hang of the programs and software really quickly, programs including Blastx, Clustal Omega, DNAdiff, etc. From learning how to code to cloning plasmids, we’ve done it all in order to identify what this cyanobacterium is. During the first few weeks of the program, we cultured the cyanobacterium in BG11 for 21 days in four different levels of salinity (25-100%). Cyanobacterium is known to be a freshwater bacteria and it was surprising to see it grow in 100% salinity. We then proceeded to manually genome annotate the strain of cyanobacterium and we were able to get results that show the whole genome sequence alignment demonstrates a 98% average identity between AB Cyanobacteria and Limnoraphis robusta. From our growing cultures we were able to practice the strategy of extracting both RNA and DNA, unfortunately extracting RNA from this particular organism is very complicated and most results were a total failure. Luckily, we were able to extract DNA. This week, after successfully extracting DNA, we proceeded to examine the concentration on the Nanodrop, and we got a really good number. This allowed us to proceed further in creating LB agar plates containing kanamycin that will be used to identify transformed cells. This whole process will allow us to create plasmids that will be shipped out for sequencing. Results are still awaiting to come back!!! Since most of the REU students, including myself, come from different states, some of us planned a weekend trip to Los Angeles, CA. The first day there, we went to the famous Santa Monica Pier. It was a dream come true, that area is so beautiful. All the shops reminded me of a calm SOHO. Later that night, we went to watch a soccer game at the Rose Bowl Stadium, where Real Madrid played against Juventus. The next day we traveled all of Hollywood, from the walk of fame, to Griffith Observatory and the Hollywood sign!!! I’ve had the time of my life these past two months!! I wish they had In-N-Out Burger back at home! Sadly it all wraps up into weeks.
0 Comments
Leave a Reply. |
Watch this space for weekly updates!Every week, one of our CSUSM NSF REU students will post their blurb, summarizing their week, and chronicling our program. AuthorWrite something about yourself. No need to be fancy, just an overview. Archives
August 2023
Categories |







 RSS Feed
RSS Feed
