 Hello everyone! My name is Trinity King, and I am a rising senior majoring in Health Science with a pre-medicine focus at Spelman College in Atlanta, Georgia. I am thrilled to have the immense pleasure of writing the first week of the 2024 summer blog. After graduation, I plan to attend medical school with the goal of becoming an Obstetrician and Gynecologist. Our program kicked off with a scavenger hunt around campus, which was a fun and engaging way to familiarize ourselves with the campus layout. We then delved into learning more about each Principal Investigator's (PI) lab and the intricate workings of their research. The next day, we were introduced to our individual labs and met the team members within each lab. The welcoming atmosphere and the emphasis on creating a family-like learning environment were truly heartwarming. I have the incredible opportunity to conduct research in Dr. Read’s lab, where we are studying coccolithophores and the process of shell calcification. I have learned how to inoculate a cell species and count cells smaller than those I am accustomed to working with. My specific project involves creating a growth curve for a particular species, which will be used for future experimentation. Today, I began the challenging process of extracting RNA from a particularly stubborn cell species. This experience will undoubtedly enhance my skills and increase my chances of successfully extracting RNA from the species I will be focusing on later. When we first arrived in San Marcos, we had a wonderful cohort dinner with Dr. Read and Dr. Lopez. It was a fantastic opportunity to get to know everyone and build a sense of community from the very start. During the week, one of the highlights was visiting the local farmer's market on Tuesday. The market boasts an array of local food vendors, beautiful greenery, unique clothing and jewelry, and the freshest fruits. It’s a vibrant and bustling place that truly captures the essence of the local culture. Over the weekend, we had an amazing trip to La Jolla, where we explored the natural caves, encountered the smelly yet fascinating sea lions, and indulged in delicious vegan waffles. We also visited a local mall, which was quite an experience. Comparing it to the malls I’m used to in North Carolina, this one had a distinct vibe – more open-air spaces, a variety of unique shops, and an overall laid-back atmosphere. It's quite different from the traditional indoor malls back home, offering a refreshing change of pace. We are excited to share more of our journey with you all in the coming weeks!
0 Comments
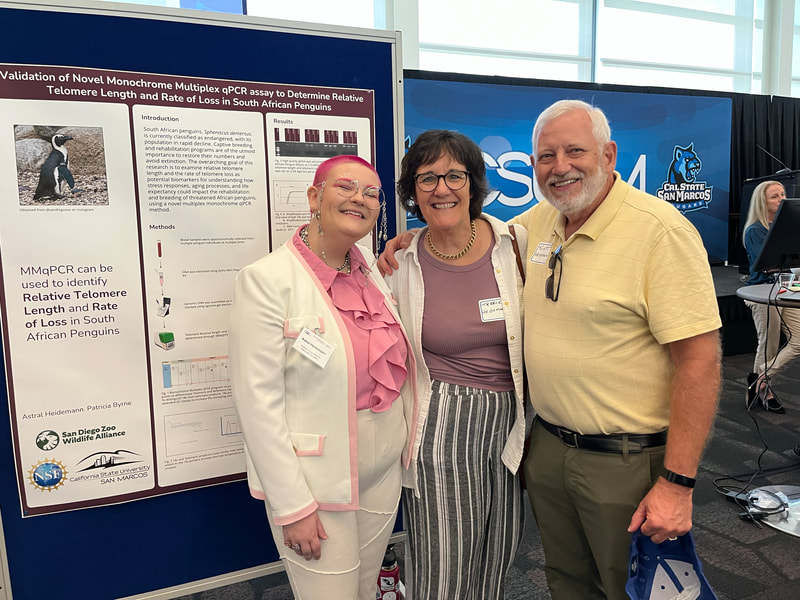
Hello!! My name is Astral Heidemann and I am a Biochemistry major! I am transferring from Palomar College to Cal State San Marcos in the fall as a rising sophomore. I am born and raised in San Diego, specifically Escondido. In highschool I had the opportunity to volunteer in a Chemistry lab at CSUSM for two years and I am very lucky to be able to come back and now work in a Biology Lab. This summer I worked with Patricia Byrnes on a project examining how stress affects relative telomere lengths and the rate of telomere loss in South African Penguin (Spheniscus demersus) samples from the San Diego Zoo. Telomeres are the protective end caps to DNA that are gradually lost during cell division. Telomere changes are closely connected to cellular aging and age-related diseases. I specifically worked on taking DNA extractions from red blood cell samples of multiple penguin individuals over multiple years, quantifying them on the Nanodrop to obtain concentration. The samples were then run on a monochrome multiplex qPCR (MMqPCR). A MMqPCR has 2 (or more) amplicons, using one fluorescence dye for both amplicons. We then used the Cq values from the MMqPCR to calculate the ∆∆Cq (the change in telomeres over years) and the fold change between years for each individual. My final poster found significant high rates of loss in two specific penguins and I am going to continue expanding data sets, including more individuals as I continue work on the project. My work with Patricia allowed me to visit the Avian Propagation Center (APC) and Beckman Center at the San Diego Zoo and Safari Park respectively. I got to see penguin egg membrane extractions and fertilization checks. I also got a tour of the APC where I saw multiple fledgling birds of varying species as well as the machines they work with. The birds were just so adorable! This final week is bittersweet; it's hard to say goodbye to the people I have become close with these last 10 weeks. It’s been a blast showing my roommates around the place where I grew up, showing them my favorite spots and restaurants, and going on fun trips with the rest of the REU such as Pride, the Barbie movie, going to the beach, seeing a Broadway show, making jewelry and many more fun outings. This week rushed by as we prepared for the poster presentation on Friday. During the poster forum it was so to walk around and see everyone else’s posters while we were setting up as well as getting the chance to show the work I have been doing this summer. We finished off by getting a final lunch together at Phò Truc Xanh which is a local family run restaurant I have been going since I was a kid and was one of the first restaurants I showed my roommates. And finally a group of us went to Escogelato Friday night to say our goodbyes to Dr. Read.
This whole summer taught me so much and I can’t wait to use all of the skills I learned this summer for the rest of my schooling and career. I am extremely excited to continue working on this project in the fall and am thankful to have been chosen for this opportunity by Professor Byrnes and to Dr. Read and Dr. Luna-Lopez for running the REU and making experiences like this possible and to my REU cohort for making this such a fun experience!  Hi everyone! My name is Janelle Mande, I am a rising senior from California State University Stanislaus, and a Biology major, minoring in Chemistry and Sociology :) This Summer, I’ve had the pleasure of working with Dr. Jane Kim, investigating molecular mechanisms that are responsible for genetic instability, or a change of length in DNA, of long CCTG repeat regions using Saccharomyces cerevisiae, or budding yeast, as our model organism. The reason why we’re so interested in these repeats is because they have the potential to cause human diseases, such as Huntington’s and Myotonic Dystrophy, yet little is known about tetranucleotide repeats in particular. My personal project involved investigating the role of transcription on CCTG DNA repeat large-scale contractions, and the most gratifying feeling I've experienced in the Kim lab is the fact that I'm actually applying what I've learned from previous semesters. My love for Biology has absolutely been reinforced with this lab, and I can’t wait to apply what I learned to future endeavors 😊 As we approach the end of the program, I realize that I have a lot to reflect on. Prior to this internship, I actually wasn’t aware that the REU program existed until the last application cycle ended. At the time, I wasn’t entirely sure if I wanted to pursue a PhD or not, but then I slowly settled on the idea of potentially pursuing one. The moment this application cycle approached, I was more than determined to apply for this Summer. Then lo and behold I got accepted into CSU San Marcos 😳 There was only ever one goal I had being a part of this program: to learn. Every day for the past 9 weeks, that’s all I’ve been doing- learning, and it amazes me that there’s still so much more I want to explore after this program ends. Research aside; however, I've done my fair share of exploring Southern California. Sprinkled across the weeks, I’ve tagged along and explored lots of places: the Salton Sea basked in heat, the cool Iron Mountain trail, and the luscious green Escondido Creek. During down time, I’ve visited Downtown San Diego, walked through the throngs of the Sunset Market at Oceanside, and explored Koreatown and Little Tokyo in Los Angeles. If you're not a local, and especially if you're not from California, I highly recommend taking the time to hang out with members of the cohort. Hanging out with folks from the cohort, if anything, has taught me how valuable it is making connections. ‘Connection’ was the staple theme I thought of when my wonderful lab partner, Maya Qaddourah, and I designed the logo for our REU sweaters. Integrated through our DNA, and in life itself, connections are important in molding who you are as a person. Needless to say, I’ll be taking a piece of everyone I’ve met in this program with me when the end comes. (And as for the folks who put effort into striking a conversation with me, you hold a special place in my heart ❤️) But alas, just as week nine draws to a close, the end feels bittersweet. It's so surreal weaving together the details of my research onto a single poster. It really puts into perspective how much hard work and effort I dedicated to the entirety of this program. I still recall the first week, and the parting words Dr. Read left us before we scattered to hunt our labs down. You really do blink, and ten weeks pass without you realizing it. To future REU cohorts, a few parting words: the hardest part is starting. Believe me, it can be tough at times. You'll naturally mess up procedures, forget to remind your PI you ran out of reagents, and the imposter syndrome will always linger. I hate to admit it, but there will even be times you feel viscerally lonely. Whether you make mistakes or feel unsatisfied with the results of your experiment, don't doubt that you're still a self-discovering scientist and learned something new. Just trust the process 🌱 Here’s my LinkedIn if you’d like to reach out to me: linkedin.com/in/jmande07  Hello from August! My name is Teresa Cajina and I am a rising senior at the University of Rochester based in Rochester, NY. I’m a declared Biochemistry major with a Certificate in Biotechnology looking forward to applying to graduate school after the end of the REU program. I grew up in a small town in New Jersey which is 5 minutes away from New York City, making me somewhat bias to the East Coast life. That being said, this program gave me the chance to visit California for the first time and there’s no other way I would’ve wanted to experience Southern California without this wholesome cohort and mentors! This week I had the opportunity to discover the Thursday night market at Oceanside (with my roommate Janelle) and adventure around Los Angeles (with Janelle and Gracie)! The first is a hands-down must for all newcomers since its accessible by the Sprinter (the light rail that has a station right next to the CSUSM University Village Apartments which is where we live during the program) and a ride only costs $2.50! Something I learned about California is that not having a car can be a major limitation in doing activities and trying out new places to eat so I highly recommend taking advantage of the Sprinter to visit Vista and Oceanside. If you do go to LA I recommend going to Porto’s bakery in Glendale first for morning breakfast since they have papa rellenas (potato balls), guava pastries and many more small eats for all taste buds priced A DOLLAR EACH! Yes, a dollar! After Porto’s my favorite spot was the Sanrio Village in Little Tokyo because it was the cutest area for the best photos and had some amazing independent-artist vendors nearby selling art, stationery, clothes and more! It is also where I had some of the best bread from Okayama Kobo bakery. Sadly, the trip was not so great at the end because the Amtrak train back to San Diego was delay by two hours due to a ‘trespasser strike’ in Anaheim so it took 5 hours in total to return to the dorms ;( Back to the research, I am currently working in the Dr. Luna lab focusing on microscopy and cell morphology of spheroids comprised of breast cancer and preadipocyte fat cells. Established in the literature, preadipocyte cells have been shown to transition between mesenchymal and epithelial states with no distinctive reasoning as to why and in the presence of breast cancer have been alluded to possibly altering the invasive properties of cancer. The lab works with both brown and white preadipocyte fat lines to observe any differences in cancer behavior. I specifically focus on the cell culture of maintaining these spheroids at different seeding densities and imaging them under 24-hour timelapses. Recently, I’ve done immunostaining with KI67 antibodies to measure proliferation within these spheroids as another means of finding factors that may influence cell structure and communication. We have two cancer lines in the lab currently called MDA-MD-231-ECADH-GFP and MCF-7-GFP which allow us to explore cancer that is not only triple negative but also fluoresce the e-cadherin protein emphasizing increased cell-cell membrane communication. I’ve had an incredible experience developing the project under Dr. Luna’s guidance that the lab was able to integrate Python programming to measure cell count and fluorescence intensity in the images making data analysis more efficient! I hope all readers have the chance to participate in a research program like the one offered at CSUSM as you’ll leave completely enriched with new friends, experiences, and life lessons! Hi :) My name is Aya Margraf, and I am a rising senior at Lewis & Clark University majoring in biology and minoring in Japanese. This summer, I have had the exciting opportunity to take part in Dr. Carlos Luna-Lopez’s breast cancer research. I have been working with two epithelial human breast cancer lines, the MCF7/GFP line, which contains ideal, in-vivo characteristics particular to the mammary epithelium, and the ECADH/GFP line, a particularly aggressive triple-negative breast cancer cell line. Along with two of my REU labmates, we have been investigating how different types of fat cells, white and brown pre-adipocytes, affect the behavior of cancer cells. For my project, I have been using biotunable frequency assembly as a cytocompatible, “bottom-up” tissue engineering technique to assemble cell spheroids, or three-dimensional cell aggregates, into tightly packed organoids. This incredibly efficient, scaffold-free method has allowed me to organize multiple assemblies of breast and fat cell organoids in a matter of seconds, mimicking native tissue to examine movement and behavior of cancer cells in the presence of host adipocytes. I have also spent alot of time using 24-hour timelapse FITC imaging, as well as challenging myself to some python coding to quantify and visually mark cell migration in various organoid organizations over time. In the midst of this all, my labmates and I have been able to properly familiarize ourselves with the routine work of cell passaging, sterilizing, technical difficulties, and laughter-inducing bits. While being blessed with some wonderful PIs, we have also had the privilege of being introduced and connected to the work of various professors and graduates students, the impressive research facilities and graduate schools in San Diego, and incredibly valuable mentor advice to navigate and enlighten us on the potential avenues up ahead. I have also acquired loads of San Diego sun! I have combed through the running trails around here, climbed in Carlsbad and Oceanside, played beach volleyball, attempted to surf on a day of no surf, paddled out in a kayak with Dr. Read at the Salton Sea, and hiked Moonlight trail to see the supermoon, a rattlesnake, and a tarantula. I also can't fail to mention that the Luna lab has never missed taco tuesdays. Never... The program is nearing its end, and gears are switching into poster presentations mode, but I have been exposed to so much this summer and have been very grateful to have been able take part in the REU at CSUSM!
Hi everyone!!!! My name is Maya Qaddourah and I am currently going into my 3rd year as a Biochemistry major here at CSUSM! My goal at the moment is to enter an MD program, hopefully somewhere in California, to eventually pursue plastic surgery/dermatology.
I have the pleasure to be working in Dr. Jane Kim’s lab, which is focused on tetranucleotide repeats, specifically CCTG nucleotides, and exploring the molecular mechanisms that may cause Myotonic Dystrophy type 2. I am currently investigating the role of Mismatch Repair (MMR) on CCTG repeat instability. I work together with my lab partner Janelle, who is also an REU student (look forward to her blog post, she has week 9!) MMR has its own set of proteins that work together to recognize small base-base mismatches, Msh2 and Msh6, and larger secondary structures, Msh2 and Msh3. Using the budding yeast Saccharomyces cerevisiae as the model organism, repeat lengths were monitored via Polymerase Chain Reactions to investigate the role of the MMR genes and their contribution to DNA repair on these CCTG repeats. We investigated the role of the DNA repair genes by knocking one out to see its effect on instability frequency. We started our week off by completing a fluctuation assay, which was able to give us the mutation rate of these contractions. We streaked 72 plates, which was not easy at all TT. My lab partner and I still had fun! We set them in the incubator to be able to count them after 72 hours. We use yeast as our model organism as it is easily manipulatable and cost-effective. My favorite part of the day is when I open the incubator to grab my projects and suddenly the lab smells like a bakery due to the yeast! Really makes me want bread….. As our week went on, we had the pleasure of visiting a few labs on the UCSD campus! We met so many awesome people and were able to tour some amazing labs. I was happy as we were able to walk into the UCSD School of Medicine, which is actually my dream school! I was really excited to see it in person and it did not disappoint. We also went down to Escondido Creek to collect some samples from the water. It was my first time visiting the creek and I was so happy to see all the nature and running water. It was so beautiful to see, especially first thing in the morning. In my free time, you will probably never catch me alone as I am always hanging out with my best friend Ileen! We do everything together like grocery shopping, working out, to watching tv. We like to go out and spend money (even though we're both broke) but that's what makes spending money even more fun. Overall, I have had so much fun in this program and I have learned more in these few weeks than I have in certain semesters! I am excited about what is yet to come in this program! Hey everyone! My name is Milana Kansky, and I am a sophomore at Palomar Community College currently pursuing my bachelor's degree in Molecular and Cell Biology. I am thrilled to be currently working in Dr. Jameson's lab and I am excited to be sharing what a week looks like as an REU student. I have been fortunate enough to have been placed on a project with a great mentor, Jake, who has been helping me get acclimated in the lab and familiar with the project. Dr. Jamesons research is focused on Gamma Delta T cells. Jake and I's project is working to try and possibly uncover a possible biomarker for Alopecia Areata. This week I have performed my own experiment using our cryostat machine to cut our Alopecia Areata (AA) skin sample into thin sheets. After we stain the skin to see specific individual cells under our IF microscope. We take pictures with the microscope and then can quantify the cells to see if there are correlations and interactions between certain cells. I feel very fortunate to be a part of Dr. Jameson's lab and I am excited to continue working with Jake and the rest of the lab members. Wednesday we were lucky enough to speak with PhD candidates that could share some wisdom to the rest of us and give us tips and tricks for grad school. Their advice was real and authentic which was refreshing to hear. On Thursday Dr. Read invited the REU cohort to join her lab in picking up some samples from the Salton Sea. It was definitely an experience I will not forget. We were hit with 104-degree weather, quicksand, and a very unpleasant smell. Despite all the heat we joked and had a great time picking up the samples and enjoyed the beautiful lake. On our way back we stopped in Julian for some delicious pie and ice-cream. When I’m not working in the lab I like to go to the beach with friends and come back to my apartment and cook dinner. This program has been a joy to be a part of and I am so grateful to be a part of it. <3  Hi! My name is Alexis Anike, and I am a senior Biology major on the Pre-Med track. I attend Howard University in Washington, D.C., but I was born in Minnesota. Post-graduation, I intend to travel abroad before pursuing an MD-PHD! This summer I am working in Dr. Read’s lab conducting research on Microcoleus Anatoxicus. Microcoleus Anatoxicus is a toxic alga that forms blooms or thick mats in freshwater systems. These toxic blooms have led to numerous canine deaths globally. I am currently attempting to extract its RNA. RNA, or ribonucleic acid, is the messenger molecule that converts the information stored in DNA into functional proteins. If I am able to extract the RNA, I will hopefully be able to determine what genes are involved in producing these toxins. It’s been a little bit of a trial and error process, as Microcoleus Anatoxicus has a very thick sheath of polysaccharides and lipids, making it hard to extract RNA from it. Outside of the lab, I have had the chance to explore California for the first time. This week, the entire REU group celebrated Juneteenth in downtown San Diego. We went on a scavenger hunt for different murals around the city and even had the chance to check out a few black-owned businesses. If you want some really good coffee and pastries, head to The Mental Bar in San Diego. We ended the day at Moonlight Beach, where we played volleyball and had a bonfire. That Wednesday, we went on a field trip to the Scripps Institution of Oceanography. It was super cool to hear about oceanography in California. They even took us out onto their pier! I ended the week with a live music festival and some thrifting with my roommates. I’m feeling a little homesick, but I'm also happy to be exploring the area! I'm glad to be part of this amazing program! Hello everyone! I hope you're all doing well. My name is Henry Basila, and I'm thrilled to share with you some exciting updates from my work in Dr. Read's lab at Cal State San Marcos. As a senior majoring in Biotechnology, I feel incredibly fortunate to be a part of this lab, as the skills, experience, and knowledge I gain here will undoubtedly pave the way for my future endeavors, particularly in pursuing a Ph.D. in cellular biology.
The past week has been truly remarkable in terms of progress and personal growth. As I delved deeper into my research project, which focuses on identifying the genes involved in the calcification process of coccolithophores, I found myself becoming more immersed and gaining a comprehensive understanding of the materials and methods within this field. Now, let me give you a brief overview of my project. Coccolithophores, fascinating unicellular algae, play a crucial role in the marine ecosystem by producing intricate calcium carbonate shells in various shapes and sizes. With over 200 different species, these organisms have an ancient lineage dating back over 200 million years. To initiate my project, I first need to determine the precise timing of shell production after a period of inoculation. This involves creating a growth curve to identify the late log phase, during which coccolithophores exhibit the highest rate of reproduction. These shells reflect light, and in order to view this calcification, I will be using a polarized light microscope to see the light reflected from their shells. Once I see calcification I will then be able to use an acid to break down the shell, and then extract rna from said decalcified algae. From there I will be doing a transcriptomic comparative analysis in order to view the difference between genes of calcified and non calcifying coccolithophores. Everyone came together to build a puzzle, either physically, or morally. After all of our hard work, Dr. Read rewarded us with some amazing crepes (what can't she do?)! Outside of the lab, I spent a lot of time with my new boyfriend, Stephan. We cooked meals together, exercised, and went to a baseball game. To end the week, I welcomed transfer students to the school as a representative of the Biotechnology club. I am very excited to spend more time in the lab, and with all of you amazing individuals!  Greetings, everyone! My name is Stephan Balensi, and I am currently an undergraduate student pursuing a degree in Business Management at San Diego State University. I am thrilled to share that I will be graduating from SDSU next year, after which I plan to embark on a Pre-Med Post Baccalaureate Program at CSU San Marcos. This program will enable me to fulfill all the prerequisites necessary for medical school. Around six months ago, my pre-med advisor introduced me to this program, emphasizing its potential to provide valuable research experience and a glimpse into the inner workings of cutting-edge medicine and treatments, even before they reach the public. 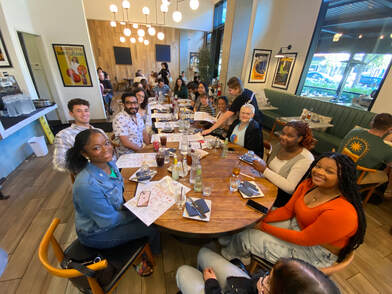 Having been born and raised in San Diego and currently residing in Chula Vista, I had the convenience of a short 40-minute drive to the San Marcos Dorms where I moved in on the first day of the program. Henry, my roommate, generously assisted me in settling into our apartment. Later, the entire REU group gathered to enjoy a delightful dinner at Buona Forchetta, an Italian restaurant. It was a wonderful opportunity to meet everyone and savor some amazing food.  The first week proved to be an incredible experience. Despite being surrounded by exceptionally bright individuals, I never felt out of place. The lab and its members welcomed me warmly, alleviating any concerns I had about diving into the extensive research that awaits us this summer. It was evident that making mistakes is part of the learning process, and the initial weeks of the program were intentionally designed to help us become comfortable with the techniques and protocols we will be using. I consider myself fortunate to have been assigned to Dr. Read's lab, which focuses on extensive algae research. During my interview with her, I mentioned that my parents own a skincare spa and produce skincare products. I also expressed my interest in potentially specializing as a dermatologist after medical school. Dr. Read thoughtfully assigned me to a project that aligns with my future aspirations!  This particular project involves culturing various Haptophyte Algae and extracting their Alkenones (lipids). The process is quite involved, including freezing the samples with liquid nitrogen, centrifuging tubes, rigorous vortexing, and intricate chemistry. Fortunately, our mentor, Hossein, is incredibly kind, knowledgeable, and patient, which significantly eased the initial apprehension I had about the protocol. Once we successfully extract the lipids, we subject them to GC-MS analysis—a combination of gas chromatography and mass spectrometry—to identify the different substances within our samples accurately. This analysis will confirm the efficiency and accuracy of our lipid extraction, as well as provide valuable insights into the composition of our samples.  With these lipids, I will be conducting tests to explore their potential as a new sunblock. In a controlled environment, I will expose pig ears to UV light after topically applying our lipids. I will also compare the results with pig ears treated with regular sunscreen and those left untreated.  Outside of the lab, we have the opportunity to embark on exciting field trips that offer a broader perspective on the industry. During this first week, we visited Thermo Fisher, where we were given an extensive tour of their state-of-the-art machinery and facilities. Additionally, we had the privilege of engaging in a conference with their esteemed faculty, where we could inquire about their journey and positions. During my leisure time, I enjoy going to the gym with fellow students and preparing healthy meals together afterward. I also take the opportunity to explore the breathtaking campus, surrounded by stunning mountains, through runs and bike rides. All in all, the first week has been nothing short of wonderful, and I couldn't be happier with the experiences and opportunities that lie ahead! |
Watch this space for weekly updates!Every week, one of our CSUSM NSF REU students will post their blurb, summarizing their week, and chronicling our program. AuthorWrite something about yourself. No need to be fancy, just an overview. Archives
August 2023
Categories |
























 RSS Feed
RSS Feed
