In 2019, Dr. Read and Dr. Hristrova went to the Anza Borrego Desert in California and discovered a desiccated stream in the San Felipe Creek of the Anza Borrego Desert. There they collected samples of desiccated cyanobacteria. Clarivel (my lab partner) and I were given the project to identify what species of cyanobacteria it is.
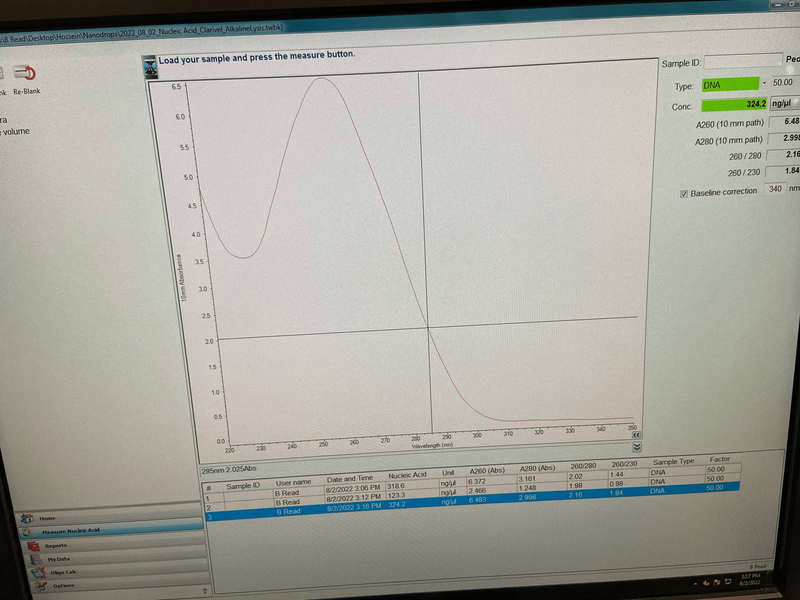
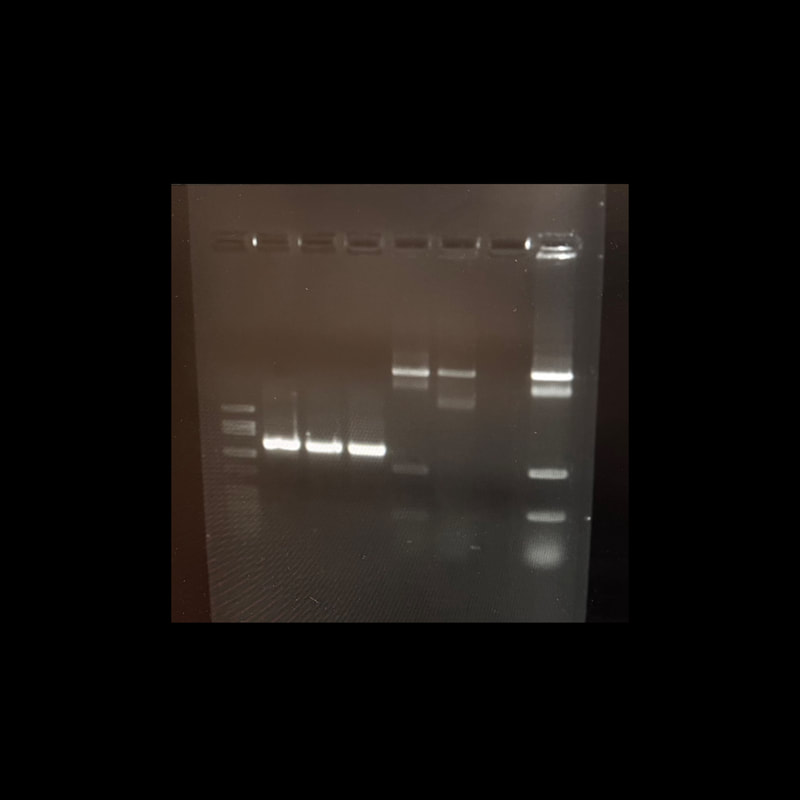
For the past 8 weeks, our initial goal for our project is to identify the unknown cyanobacterium from the Anza Borrego San Felipe Creek with whole genome sequencing and then to interrogate its phylogenetic history. During this time I have learned a lot of bioinformatic tools and techniques. I’ve also heard the horrors and tragedies of bioinformatics, and how important it is to maintain detailed notes of the purpose and methods used in computational analysis. With the help of Dr. Read, I was able to quickly get the hang of the programs and software really quickly, programs including Blastx, Clustal Omega, DNAdiff, etc. From learning how to code to cloning plasmids, we’ve done it all in order to identify what this cyanobacterium is. During the first few weeks of the program, we cultured the cyanobacterium in BG11 for 21 days in four different levels of salinity (25-100%). Cyanobacterium is known to be a freshwater bacteria and it was surprising to see it grow in 100% salinity. We then proceeded to manually genome annotate the strain of cyanobacterium and we were able to get results that show the whole genome sequence alignment demonstrates a 98% average identity between AB Cyanobacteria and Limnoraphis robusta. From our growing cultures we were able to practice the strategy of extracting both RNA and DNA, unfortunately extracting RNA from this particular organism is very complicated and most results were a total failure. Luckily, we were able to extract DNA. This week, after successfully extracting DNA, we proceeded to examine the concentration on the Nanodrop, and we got a really good number. This allowed us to proceed further in creating LB agar plates containing kanamycin that will be used to identify transformed cells. This whole process will allow us to create plasmids that will be shipped out for sequencing. Results are still awaiting to come back!!! Since most of the REU students, including myself, come from different states, some of us planned a weekend trip to Los Angeles, CA. The first day there, we went to the famous Santa Monica Pier. It was a dream come true, that area is so beautiful. All the shops reminded me of a calm SOHO. Later that night, we went to watch a soccer game at the Rose Bowl Stadium, where Real Madrid played against Juventus. The next day we traveled all of Hollywood, from the walk of fame, to Griffith Observatory and the Hollywood sign!!! I’ve had the time of my life these past two months!! I wish they had In-N-Out Burger back at home! Sadly it all wraps up into weeks.
0 Comments
 Hello everyone, my name is Marvin Onwukwe, and I am a rising senior at the University of Maryland Baltimore County (UMBC) majoring in Biological Sciences. I have been working under Dr. Jane Kim’s lab, investigating the genetic instability of microsatellite regions of DNA using yeast. The project I had the pleasure of working on involved performing knockouts of genes in the highly manipulable genome of yeast to investigate mutation rates of repeat sequences to gain a better understanding of the currently incurable genetic illness known as Myotonic Dystrophy type 2 and potential therapeutic treatment. Among the myriad of lab skills Dr. Kim and her work had exposed me to, I had the opportunity to develop into a more independent researcher and academic in general having had the experience of working on a project for the first time. I want to emphasize being kind to yourself especially in the first few weeks. Experiments may go wrong, equipment may be unfamiliar, and hiccups in general do happen. That is only a reflection of the fact that you are challenging yourself in a new environment. Even though perfection is ideal, don’t let it be the gateway that allows you to enjoy yourself and what you’re learning in the lab. I am excited to write to you about week 7 which happened to be the week of my 21st Birthday. That week, we were treated to a tour of the Scripps Institution of Oceanography, a world class facility dedicated to advance our understanding of the earth system and train the next generation of scientific and environmental leaders. As a department of UC San Diego, Scripps Institution of Oceanography is one of the oldest, largest, and most important centers for ocean, earth, and atmospheric research. The tour had explained how they make accurate oceanic simulations in large-scale, cleverly controlled environments among many other features. Not only are we exposed to world class facilities and quality preparation for graduate education, in California you genuinely have a higher prevalence/chance of meeting some of your favorite celebrities. On the weekend of my birthday, I had gone to downtown San Diego to celebrate, and I was elated to meet Brian Posehn on the sidewalk by chance. For any of those who watched the show Big Bang Theory, he was the geologist at Cal Tech. I would like to mention that the following week (week 8), I had the opportunity to travel to Los Angeles, which is a mere 40-minute Sprinter train from the CSUSM campus to the Oceanside Amtrak ($2.50) and a two-hour Amtrak from there to LAX union station. As part of the McNair scholars’ program at my University in Maryland I was able to join them and meet the CSUSM McNair scholars at our national conference held at the University of California, Los Angeles (UCLA) where I presented research and had tremendous networking opportunities and graduate school prep similar to the standard given at this REU experience. I couldn’t recommend this opportunity more! Here is my LinkedIn: Marvin Onwukwe Hi everyone! I am Chloe La Prairie, and I am a senior at Millikin University in Decatur, Illinois studying Biology with a pre-med concentration. This summer I have been working in Dr. Jameson’s lab conducting Neuroimmunology research! I have been studying the relationship between Gamma Delta T Cells and neurons in wound repair. During my six weeks in the lab so far, I have learned an immense amount while doing experiments. This week I conducted multiple ear peel experiments, which is where we peel apart the epidermis from the dermis of mouse ears and stain with different antibodies to look at the relationship between Gamma Delta T Cells and the neurons through the immunofluorescent microscope. We do this with wounded and non-wounded mice to examine how Gamma Delta T Cells affect wound repair. In addition to ear peels, I have also learned how to do cross sectioning cut staining and culture cell lines. When I am not in the lab, I am spending my time studying for the Medical College Admission Test which I am taking at the end of July and finishing up my applications to medical school. This week the REU students had the opportunity to go down to San Diego State University and tour several research labs. It was very interesting to see how different labs work and what other kinds of research are currently being conducted. On Saturday, we all went over to Dr. Reads house for dinner which was so much fun. Dr. Read even had homemade ice cream and crepes for us! On the weekends, I look forward to spending time with the others in the REU program, and I’ve enjoyed spending most of my weekends at the beach or the pool. I also enjoy spending time with my lab mates outside of lab, we have met up for Pizookie Nights at BJs Restaurant and walked down to the farmers market together. I look forward to learning more skills in lab that I can then take back with me to my own university where I will continue to conduct research. I am very grateful for the opportunity the REU program was given me this summer and will be sad when the summer ends. I can confidently say this is one of the best summers I have ever had! This week I learned how to digest Malpighian tubules from Trichoplusia ni (a species of caterpillars) while waiting for our purchased insect medium to arrive. These tubules help scientists understand Trichoplusia ni osmoregulatory and excretory system. Along with digestion, I began to prepare for cell culture, which involved autoclaving my materials and sterilizing an area in the lab. Cell culture in this lab has never been done before so I am a bit nervous, but excited to be a part of this process!
After our field work, we visited Julian to feast on their famous pies before heading back to campus. Later that day, I attended a Mentor Fest with other interns where I had the opportunity to listen and ask questions related to the science field and potential career paths. Although many of them are not in job positions I am interested in, it was fulfilling to hear their life journeys and the endless advice they had to offer. Gaining perspectives of people in the working field was such a motivating and reassuring experience. I am truly grateful for the opportunity as it helped me decide on my plans post undergraduate school.
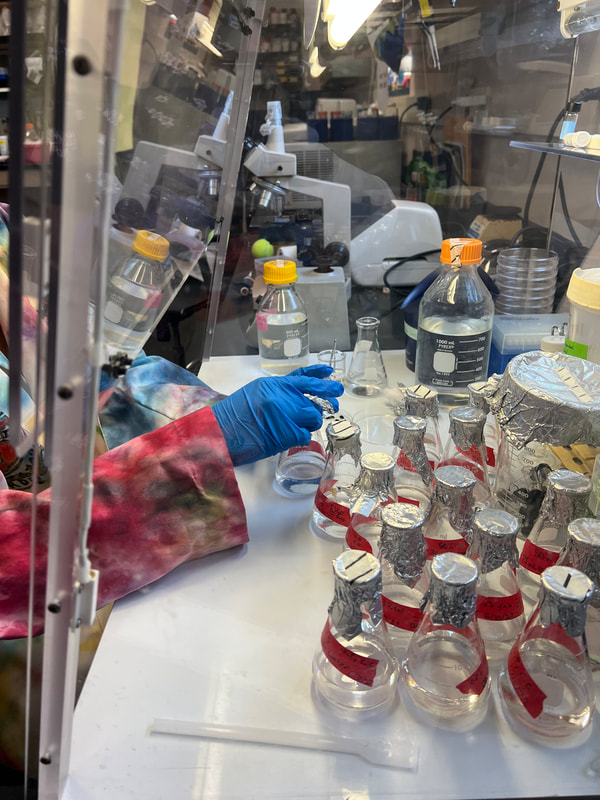
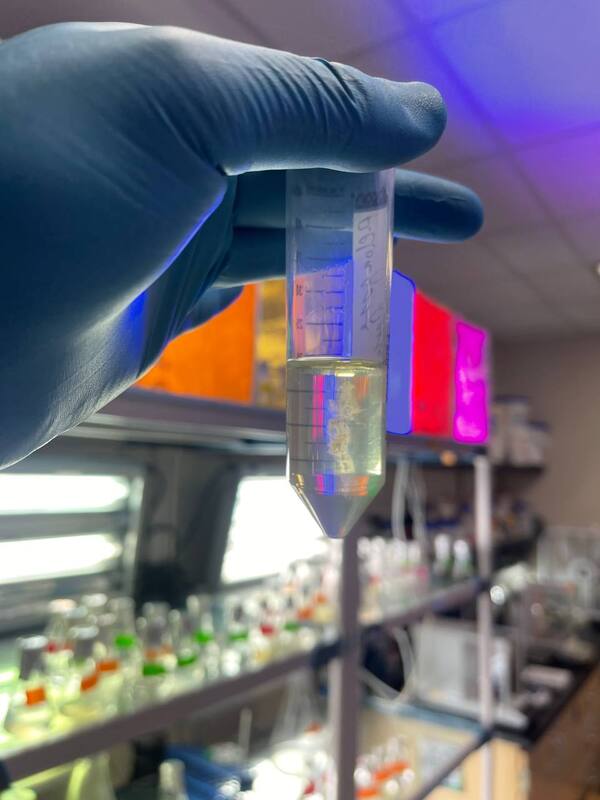
It has been great getting to hang out with the amazing REU and Summer Scholar interns. I can't believe we are halfway through the internship already! There is still a lot I want to learn at CSUSM and places I want to adventure to with the other interns. We have a lot of ideas that I hope we get to do before the end of summer. But I am going to make the most out it and enjoy it all <3 Hey y’all! My name is Jenna Archambeau and I am a rising senior at American University in Washington, DC. I study biochemistry and statistics, which I have been able to apply here at CSUSM for next-generation sequencing and bioinformatics! For this summer, I have been working with Dr. Elinne Becket in her microbial genomics lab. We work with coastal water metagenomes to get a better understanding of horizontal gene transfers in response to tetracycline, which we hypothesize to have an impact on community antibiotic resistance. This week I have been assisting others in the lab on their projects. Last week, Ella (another REU student in Dr. Becket’s lab) and I troubleshooted the protocol for our RNA Library Prep because the samples contain RNA from both eukaryotes and prokaryotes. We are hoping to isolate our mRNA from bacteria, so we need to eliminate ribosomal RNA from the eukaryotes and prokaryotes. Depleting rRNA from both sources turned out to be more difficult than expected, so we had to consult with various biotech companies to figure out our best option moving forward. It was nerve racking to make all of the technical support calls; however, it was rewarding to get more pieces to the puzzle. While waiting for our supplies to come in, I have been helping other students in the lab with their projects. On Tuesday, we went out to Scripps Pier to take ocean water samples for one of our graduate students. I enjoy field work because seeing the environment we are studying makes our lab goals more tangible. The rest of the week I have been developing both dry lab and wet lab skills by working on code and DNA extraction. Though I have used R and CommandLine previously, it has been a welcome challenge to explore new pipelines and applications of these resources. It was fascinating to learn how many different ways you can manipulate the code to get results that are relevant and significant toward your research question. Lastly, I conducted a DNA extraction experiment from Vibrio bacterial cells for the first time. I had a lot of fun with the experiment, as you can tell from the picture. Outside of the lab, I have made a lot of great memories with my fellow REU students. This weekend, we celebrated our long weekend for the Fourth of July together by hanging out by the pool, going to the beach, and playing laser tag. I’m hoping to explore more beaches (maybe learn surfing?) and experience everything Southern California has to offer during my time here. It’s been amazing to connect with others, both inside and outside of the lab, and I look forward to what’s next to come in the next six weeks!! 
REU DR. Read’s lab This week I successfully extracted genomic DNA and RNA from the cultures I grew up in the past two weeks. There are some practical goals in my research project. One is performing all the processes of DNA and RNA extraction sterilely to avoid any contaminations. The second one is extracting genomic DNA. To do so, DNA should not be sheared in anyway. With that being said I was not able to use pipetting methods to transfer my DNA content which can be challenging sometimes. Also, I had to use a new method to pellet the algae instead of centrifuging it. This week, Dr. Read; my mentor, decided to use a filtration method instead of spinning the cells. It was meow first-time using filtration method to get the cells out of their cultures. Although it took about two hours to filter all the cells, I am so happy that the method worked and I could extract the DNA and RNA from my samples. The ultimate goal of this project I am working on is to understand the algae’s characteristics in genomic level. I also started to help Dr. Read with another project which I found very interesting. I started to inject gas chromatography and mass spectrometry samples in GC-MS machine. The samples are from different specious such as Emiliania huxleyi. In this project we would like to identify the components of Emiliania huxleyi’s lipids at a molecular level. I started to learn how to work with the machine and the software program which is being used to operate the machine. I would like to thank my great mentor Dr. Read for conveying her expertise to me passionately and being willing to teach and help all the time. Bioinformatics In our Wednesday bioinformatic meeting I learn a lot in many ways. This week we started to use new tools in Kbase website such as Kaijo. We also started to analyze the FASTQC results from the previous weeks. I believe our bioinformatic meetings are very important for one main reason and that is the necessity of bioinformatics in our filed of work. We pretty much know how to practically generate the data that we think can be helpful for our research project. However, understanding the generated data and making biological sense out of them is what bioinformatics does, and it can be quite challenging. This week we have to analyze some of the generated data with different tools in Kbase website and present them in our next week bioinformatics meeting. Next week I will run agarose gels for my extracted DNA and RNA and I can’t wait to see what the results are. I am also looking forward to present the table that we generated for results section of our manuscript in our bioinformatics meeting. Activities Beside work which was so exciting, I had a great week in terms of the fun activities that we all did. First of all, I celebrated my birthday with my REU colleagues at our pool. It was a great party and we all enjoyed the pool in on a sunny Saturday. Before that some of my REU friends distracted me for a day to arrange a surprise birthday which they did a great job doing that. In the middle of the week I got a chance to visit Thermo Fisher’s Carlsbad site for the second time. This time I went with a group of CSUSM students who are mostly looking for job immidiently or sometime soon. It was great meeting both graduated and senior peers in a different setting than collage. Next week are going to a field trip to Anza-Borrego desert state park, which I’ve been waiting for a long time. Hi y’all! My name is Cristina Velasquez I am a rising junior and will be transferring here to CSUSM this Fall from Palomar Community College! I’ll be majoring in Biological Sciences with an emphasis in Physiology. I live in northern San Diego County in a little town called Pauma Valley (orange and avocado tree central). I am very grateful to have been accepted into this program! It has been nothing but fun so far! I joined Dr. Jameson’s lab in January through a program offered at my old college. I was observing and learning the basics of the lab and did some experiments with my lab mates and once this program started I began to do A LOT more hands on work. I am now on the alopecia areata project. Here I am on a team of 3 and we are looking at the effect gamma delta t-cells have on the hair follicle. So far I have done a couple ear peels which is a procedure where we peel apart the epidermis from the dermis of mouse ears. We then stain the peels with our antibodies of interest and look at our slides through the immunofluorescent microscope in the IF room which is a room with a dim red light to lower the light exposure since our antibodies are light sensitive. After spending hours in that room I can definitely say my eyes take a couple minutes to adjust to natural light! I have also been using the microscope on my own when doing cut staining which is a task you’d think would be easy…it’s just a microscope…nope! There’s lots of little details that need to be accounted for and it’s pretty delicate. I’ve learned how to use the crysostat which is a machine that allows us to cut our frozen skin sections and see what they look like under the microscope. It’s a “glorified deli slicer”! We mainly use this machine for cut stainings. I can’t wait to learn more skills that will make me a better researcher and eventually complete experiments all on my own by the end of the summer! Outside of the lab our REU cohort attends seminars on Wednesdays, and this week was an introductory lecture on Next Generation Sequencing given by Dr. Becket. We then had some guest speakers come in from the San Diego Safari Park Research Center and they talked about Sequencing the Genome of the Northern White Rhino. It was a very interesting talk! The Jameson lab has journal clubs each week so we all meet at a local business for a snack, coffee, or lunch and talk about the research paper for that week in a round table like discussion. This week we went to Copa Vida and talked about a paper on how dysfunctional gamma delta t-cells contribute to impaired keratinocyte homeostasis in mouse models. The day before journal club Chloe (who is a fellow REU student and is also in the Jameson lab) and I went to Tamarack Beach in Carlsbad to read the paper. It was a fun afternoon of snacks and scientific reading! Friday night after going out for dinner all the REU students hung out in our dorm and we had a mini dance party and ate my left over sushi and edamame! Saturday came and we went to Ponto Beach in Carlsbad for half the day. It was the first good beach day in a long time so it was insanely packed but it was still very fun! I went home for Fathers Day on Sunday and had some yummy food with my family! On a side note…I bought a surfboard this week lol! I am really looking forward to using it soon! Week 2 was amazing! I can’t wait for the next 8 weeks! Time is already flying by but I am enjoying and soaking up every minute of it!  Hello! I am Christian Ihenyen, a rising senior at Howard University majoring in Biology. I am originally from Durham , NC but I'm in love with this Southern California weather. Coming into the program I was excited and curious because I had never been to the west coast before. After meeting other participants, I knew this was the program for me. My counterparts were very welcoming and willing to socialize. The first week was nothing like I would have expected. My lab made me feel very welcomed. Sometimes when you come to a space that you are not familiar with it can be really nerve wracking when it comes to not wanting to make mistakes. One thing I can say is that I'm learning something new everyday. Thanks to Dr. Read, she has a good way of setting things clear and being patient with the students. In the lab, I'm investigating the effects of salinity on anatoxin production. Right now I'm preparing my RNA extractions and I've learned so much already, for example, extraction is a bit different from DNA extraction, you have to be very fast and clean because RNA molecules degrade easily. On Tuesday we went to a delicious taco spot and I ordered almost everything on the menu. Our first day, we visited a big scientific corporation called Thermo fisher. Thermo Fisher is a very big building right in the beautiful city of Carlsbad. I was really excited to listen the panel because most of them weren't STEM majors but they had careers in science. Hearing that made me more comfortable with not having to follow a certain path because of my major. I even got the chance to introduce myself to one of the panelists who has a career in data analyst therefore I made sure to get his contact information. The fun didn't stop there though. On Saturday we went to Torrey Pines to hike and explore the beaches well. Then we went to pacific beach to celebrate fellow REU intern, Jenna's 21st birthday and this taco spot called The Taco Stand. It was definitely a success, there was some random man who even offered to pay for our McDonalds . That Sunday I took time to myself to whine down and prepare for the lab the next day. So far, the first week has been filled with great memories and the beginnings in my research career. I can’t wait for what lies ahead!
Hello everyone. My name is Isabelle Massaro and like Antolette, I am fortunate to be one of the local California State University, San Marcos (CSUSM) students accepted into the NFS REU Research Program this summer. I am a senior here at CSUSM (with only 1 more semester left!) and am majoring in Molecular and Cellular Biology with a minor in Chemistry. As I near the end of my bachelor's degree, I plan to continue my education to pursue my passion in plant biology research. Since I joined Dr. Escobar's molecular plant biology lab, I have gained hands-on experience working with plants. My project in the Escobar lab focuses on working with the model plant, Arabidopsis thaliana, to help characterize class III glutaredoxins (GRXs) and identify their role in plant growth and development. GRXs are small oxidoreductase enzymes that reduce disulfide bonds in target proteins. The interesting thing about class III GRXs, is that they are exclusively found in land plants, but most remain functionally uncharacterized. To help identify the biological function of these genes, we have created different transgenic plant lines that either overproduce class III glutaredoxins, or have knockout mutations in class III glutaredoxin genes. For instance, this summer I have been growing two knockout lines that have five GRX genes (AtGRXS3/4/5/7/8) inactivated and two overexpression lines that constitutively overexpresses AtGRXS8. We previously found that this cluster of class III GRXs are significantly upregulated by nitrogen in the soil, specifically in the form of nitrate. This is important because nitrogen is essential for plant growth and development and without nitrogen, plants can not produce DNA, RNA, proteins, or chlorophyll. This summer, my main focus has been on extracting RNA from hydroponically grown transgenic plants with the end goal of sequencing the whole transcriptome using Next-Generation Sequencing (NGS). This will help identify how our GRXs influence patterns of gene expression in plants provided with nitrate as a nitrogen source. The start of the summer involved lots of tissue grinding, RNA extractions, gels for quality control, and cDNA synthesis. This week was especially exciting because we were finally able to start performing Real-Time PCR runs on all of the samples. From these results, we can identify differences in gene expression between our transgenic lines and the wildtype for some specific nitrate transporter genes in roots and shoot growth genes. These results will also help us identify the best samples to send for RNA sequencing. In addition to the molecular work that I have been doing, we have also been growing lots of plants on soil for phenotyping. This allows us to better understand how GRXs affect overall plant growth. For instance, our overexpression plants have significantly smaller shoots compared to the wildtype, which is why we hypothesize that these GRXs genes are negative regulators for shoot growth. As for phenotyping, I have been tracking flower production, weighing total shoot biomass, and imaging pavement cells. And within the next couple of weeks, we plan to complete total nitrate and protein content analyses to better understand how efficiently our transgenic plants store and utilize nitrogen. Overall this allows us to help characterize these class III GRXs and their role in plant growth and development. At the end of the week, we were finally able to go tide pooling! Low tide this summer has been a little tricky to plan around, since it was always very early in the morning. That did not deter us though, and a group of us got up at 4:00 AM to venture out to the coast. We ending up going to Sunset Cliffs National Park and it was well worth waking up so early. This was probably one of the best tide pooling experiences I have had. We found so many neat creatures that I have never seen before. Immediately when we got there, we found an octopus hiding within the algae, which was a great start to our early tide pool adventure. After that, we also found an eel, a lobster, a beautiful red Hopkins’ rose nudibranch, a delicate brittle star, and a ginormous sea hare. And there were so many crabs, shrimps, and little isopods scurrying everywhere. I am very grateful to be a part of such an incredible cohort this year. This summer has definitely been an amazing experience and I want to thank Dr. Sethuraman and Dr. Read for running an excellent program during these challenging times. Good luck with everyone's research project as we come to the home stretch.
|
Watch this space for weekly updates!Every week, one of our CSUSM NSF REU students will post their blurb, summarizing their week, and chronicling our program. AuthorWrite something about yourself. No need to be fancy, just an overview. Archives
August 2023
Categories |





















































 RSS Feed
RSS Feed
